mRNA Splicing - mRNA post-transcriptional processing/modifications - What is alternative splicing?
What is Splicing?
General Definition and Overview
- Splicing is the process of removing introns and, in some cases, exons from pre-mRNA (PMRNA) during or after transcription.
- The video will be divided into three parts: what splicing is, how it occurs, and why it is necessary.
Structure of Pre-mRNA
- A simple PMRNA structure consists of two exons (E1 and E2) separated by an intron (I). The focus will be on the intron syntax.
- Intron syntax includes specific nucleotide sequences at both ends: a guanine-uracil (GU) pair at the 5' splice site and adenine-rich sequences towards the end.
Intron Syntax Details
Key Components of Intron Syntax
- The five prime splice site has a consistent presence of GU nucleotides; this is crucial for splicing initiation.
- The branch point within the intron contains adenines, followed by a pyrimidine track rich in U's and C's leading to the three prime splice site enriched with adenine and guanine.
Types of Introns
- Approximately 90% of PMRNAs contain U2 type introns characterized by specific splice sites; remaining 10% are U12 type with different nucleotide compositions at their splice sites.
Types of Introns Beyond Pre-mRNA
Classification of Introns
- There are various types of introns including nuclear pre-mRNA (U2 & U12), organelle-specific introns, and bacterial introns which can be self-splicing or require endonucleases for removal.
- Self-splicing introns act as ribozymes that do not need proteins for their removal; they are found in mRNAs, tRNAs, and rRNAs across different organisms.
Spliceosome Functionality
Role in Splicing Process
- Focus will remain on splicosome-dependent U2 type introns where splicing begins once the splicosome proteins recognize the intron syntax.
Composition of Spliceosomal Proteins
- Small nuclear ribonucleoproteins (snRNPs), composed of small nuclear RNA (snRNA) and protein components, play a critical role in recognizing these signals during splicing processes. Major snRNP types include U1, U2, U4, U5, and U6.
Mechanism of Splicing
How Splicing Occurs
- The process involves complex interactions among snRNP components that form secondary structures essential for recognizing splice sites within the PMRNA strand containing a U2 type intron.
Splicing Mechanism Overview
Recognition Stage of Splicing
- The recognition begins at the five-prime splice site where U1 snRNP pairs with GU in pre-mRNA, marking the initial step in splicing.
- SR proteins, characterized by serine and arginine repeats, stabilize U1 snRNP binding to RNA and assist in splicing regulation.
- The branch point adenine is recognized by splicing factor 1 (SF1), while U2 AF protein dimer binds to the three-prime splice site, enhancing recognition through cooperative binding.
- This recognition occurs in a looped conformation due to long introns, facilitating interactions between U1, SF1, and U2 AF proteins.
- The early complex forms as SR proteins help bring exons closer together; this complex is also known as a stable or committed complex.
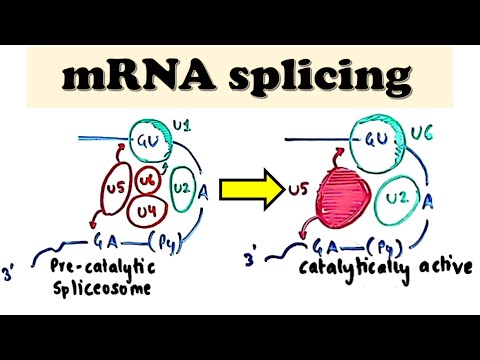
Formation of Pre-catalytic Complexes
- After the early complex assembles, U2 snRNP replaces SF1 and U2 AF proteins at the branch point, forming the A complex while maintaining U1 at the five-prime splice site.
- The B1 complex emerges when U5, U4, and U6 join; here, U5 bridges exons while replacing SR proteins' function within the assembly.
- To activate catalysis, both U1 and U4 are removed from this assembly; thus creating a B2 complex that is catalytically active for splicing reactions.
Catalytic Steps of Splicing
- The first transesterification reaction occurs when hydroxyl from adenine attacks phosphate at the five-prime splice site; this cleaves off exon from intron.
- As exons are cut free from intron lariat structure remains bound to snRNP complexes; this leads to formation of catalytic one complex.
- In the second transesterification step, upstream exon attacks three-prime splice site leading to ligation of two exons while releasing intron lariat structure.
- The resulting structure post-ligation shows two joined exons with an attached lariat-shaped intron still associated with snRNP complexes.
- Special RNA helicases facilitate removal of SF1 and other components during these steps which are crucial for successful splicing.
Detailed Mechanism Insights
- Examination reveals phosphodiester linkages between G and U at five-prime splice site alongside sugar-phosphate connections around branch point adenine during transesterification steps.
- First transesterification involves attack by two-prime hydroxyl on guanosine phosphate leading to formation of looped intron lariat structure observed in catalytic one complex.
- Second step sees upstream exon’s free hydroxyl group attacking three-prime splice site phosphate which releases intron lariat allowing mRNA formation through exon joining.
Understanding Splicing Mechanisms
The Basics of Splicing
- Splicing is crucial for removing introns and joining exons in mRNA, which is essential for producing functional proteins.
- Canonical splicing involves the independent removal of introns and the joining of exons, allowing for various combinations to form different mRNAs.
- The removal of introns prevents the production of defective proteins since introns are non-coding regions. This process enhances protein diversity through alternative splicing.
Alternative Splicing Explained
- Alternative splicing allows a single gene to produce multiple RNA isoforms by varying exon combinations, significantly increasing protein diversity within cells.
- To achieve specific exon arrangements (e.g., E1 and E3), certain proteins must prevent other exons from participating in splicing, showcasing the complexity of this regulatory mechanism.
Trans Splicing in Worms
- In model organisms like worms, trans splicing occurs where exons from two different RNAs are joined together, involving unique elements called outrons.
- The splice leader RNA (SLRNA), containing a short exon and an intron, plays a key role in trans splicing by transferring important features like the 5' cap to stabilize mRNA.
Understanding Outrons
- During trans splicing, the SLRNA's branch point interacts with the PMRNA's splice site to replace its 5' UTR with that from SLRNA, enhancing stability.
- Outrons differ from traditional introns; their removal results in a branched lariat structure distinct from typical intron lariats observed during canonical splicing.
Back Splicing: A Unique Process
- Back splicing involves an exon attacking its own previous end rather than following standard forward directionality during canonical splicing.
- This type of splicing can lead to circular RNA formation and is often associated with pathological conditions rather than normal cellular processes.